2017-2020 Publications
2017-2020 Publications
2017-2020 Publications
2012-2016 Publications
Highlights
* = co-first author, † = corresponding author

Cryo-EM Structure of Caspase-8 Tandem DED Filament Reveals Assembly and Regulation mechanisms of the Death-Inducing Signaling Complex
Fu TM*, Li Y*, Li Z, Vajjhala PR, Cruz AC, Srivastava DB, DiMaio F, Penczek PA, Siegel RM, Stacey KJ, Egelman EH, Wu H†. Mol Cell (2016). PDF
Caspase-8 activation can be triggered by death receptor-mediated formation of the death-inducing signaling complex (DISC) and by the inflammasome adaptor ASC. Caspase-8 assembles with FADD at
the DISC and with ASC at the inflammasome through its tandem death effector domain (tDED)... Read More

Inflammasome-activated gasdermin D causes pyroptosis by forming membrane pores.
Inflammatory caspases (caspases 1, 4, 5 and 11) are activated in response to microbial infection and danger signals. When activated, they cleave mouse and human gasdermin D (GSDMD) after Asp276 and Asp275, respectively, to generate an N-terminal cleavage product (GSDMD-NT) that triggers inflammatory
death (pyroptosis) and release of inflammatory cytokines such as interleukin-1β1,2... Read More

The Structure and Dynamics of Higher-Order Assemblies: Amyloids , Signalosomes, and Granules
We here attempt to achieve an integrated understanding of the structure and dynamics of a number
of higher-order assemblies, including amyloids, various kinds of signalosomes, and cellular granules. We propose that the synergy between folded domains, linear motifs, and intrinsically disordered regions regulates the formation and intrinsic fuzziness of all higher-order assemblies, creating a structural... Read More

Molecular basis of caspase-1 polymerization and its inhibition by a new capping mechanism
Lu A*, Li Y*, Schmidt FI, Yin Q, Chen S, Fu TM, Tong AB, Ploegh HL, Mao Y, Wu H†. Nat Struct Mol Biol (2016). PDF
Inflammasomes are cytosolic caspase-1-activation complexes that sense intrinsic and extrinsic danger signals, and trigger inflammatory responses and pyroptotic cell death. Homotypic interactions among Pyrin domains and caspase recruitment domains (CARDs) in inflammasome-complex components mediate oligomerization into filamentous assemblies. Several cytosolic proteins... Read More

A single domain antibody fragment that recognizes the adaptor ASC defines the role of ASC domains in inflammasome assembly
Myeloid cells assemble inflammasomes in response to infection or cell damage; cytosolic sensors activate pro–caspase-1, indirectly for the most part, via the adaptors ASC and NLRC4. This leads to secretion of proinflammatory cytokines and pyroptosis. To explore complex formation under physiological conditions, we generated an alpaca single domain antibody, VHHASC, which specifically recognizes... Read More
.jpg)
Structural basis and functional role of intramembrane trimerization of the Fas/CD95 death receptor.
Fu Q*, Fu TM*, Cruz AC, Sengupta P, Thomas SK, Wang S, Siegel RM, Wu H†, Chou JJ†. Mol Cell (2016). PDF
Fas (CD95, Apo-1, or TNFRSF6) is a prototypical apoptosis-inducing death receptor in the tumor necrosis factor receptor (TNFR) superfamily. While the extracellular domains of TNFRs form trimeric complexes with their ligands and the intracellular domains engage in higher-order oligomerization, the role of the transmembrane (TM) domains is unknown. Read More
.jpg)
Molecular mechanism of V(D)J recombination captured by structures of RAG1-RAG2 synaptic complexes.
Diverse repertoires of antigen-receptor genes that result from combinatorial splicing of coding segments by V(D)J recombination are hallmarks of vertebrate immunity. The (RAG1-RAG2)2 recombinase (RAG) recognizes recombination signal sequences (RSSs) containing a heptamer, a spacer of 12 or 23 base pairs, and a nonamer (12-RSS or 23-RSS) and introduces precise breaks at RSS-coding segment junctions. Read More

Cryo-EM structure of the activated NAIP2/NLRC4 Inflammasome reveals nucleated polymerization.
Zhang L*, Chen*, Jianbin R, Wu J, Tong AB, Yin Q, Li Y, David L, Lu A, Wang WL, Marks C, Ouyang Q, Zhang X, Mao Y†, Wu H†. Science (2015). PDF
The NLR family apoptosis inhibitory proteins (NAIPs) bind conserved bacterial ligands, such as the bacterial rod protein PrgJ, and recruit NLR family CARD-containing protein 4 (NLRC4) as the inflammasome adapter to activate innate immunity. We found that the PrgJ-NAIP2-NLRC4 inflammasome is assembled... Read More

Structural Biology of Innate Immunity
Yin Q*, Fu TM*, Li J, Wu H. Annu. Rev. Immunol (2015). PDF
Innate immune responses depend on timely recognition of pathogenic or danger signals by multiple cell surface or cytoplasmic receptors and transmission of signals for proper counteractions through adaptor and effector
molecules. At the forefront of innate immunity are four major signaling pathways, including those elicited by Toll-like receptors, RIG-I-like receptors, inflammasomes, or cGAS, each with its own cellular localization, ligand
specificity, and signal relay mechanism. They collectively... Read More

SMOCs: supramolecular organizing centres that control innate immunity
Kagan JC*†, Magupalli V*, Wu H*†. Nature Rev Immunol (2014). PDF
The diverse receptor families of the innate immune system activate signal transduction pathways that are important for host defence, but common themes to explain the operation of these pathways remain undefined. In this Opinion article, we propose — on the basis of recent structural and cell biological studies — the concept of supramolecular organizing centres (SMOCs) as location-specific higher-order signalling complexes in which increased local concentrations of signalling components promote the intrinsically weak allosteric.. Read More
.jpg)
IRAK4 Dimerization and the Trans-autophosphorylation are Induced by Myddosome Assembly
Trans-autophosphorylation is among the most prevalent means of protein kinase activation, yet its molecular basis is poorly defined. In Toll-like receptor and interleukin-1 receptor signaling pathways, the kinase IRAK4 is recruited to the membrane-proximal adaptor MyD88 through death domain (DD) interactions, forming the oligomeric Myddosome and mediating NF-kB activation. Read More
.jpg)
Unified Polymerization Mechanism for the Assembly of ASC-Dependent Inflammasomes
Lu A*, Magupalli VG*, Ruan J*, Yin Q, Atianand MK, Vos MR, Schröder GF, Fitgerald KA, Wu H†, Egelman EH. Cell (2014). PDF
Inflammasomes elicit host defense inside cells by activating caspase-1 for cytokine maturation and
cell death. AIM2 and NLRP3 are representative sensor proteins in two major families of inflammasomes. The adaptor protein ASC bridges the sensor proteins and caspase-1 to form ternary... Read More
.jpg)
Structural Architecture of the CARMA1/Bcl10/MALT1 Signalosome: Nucleation-Induced Filamentous Assembly
Qiao Q*, Yang C*, Zheng C*, Fontan L, David L, Yu X, Bracken C, Rosen M, Melnick A, Egelman E, Wu H†. Mol Cell (2013). PDF
The CARMA1/Bcl10/MALT1 (CBM) signalosome mediates antigen receptor-induced NF-kB signaling to regulate multiple lymphocyte functions. While CARMA1 and Bcl10 contain caspase recruitment domains (CARDs), MALT1 is a paracaspase with structural similarity to caspases. Here we show that the reconstituted CBM... Read More

Molecular Mechanism for p202-mediated Specific Inhibition of AIM2 Inflammasome Activation
Yin Q*, Sester DP*, Tian Y*, Hsiao YS, Lu A, Cridlan JA, Sagulenko V, Thygesen SJ, Chouhey D, Hornung V, Walz T, Stacey KJ†, Wu H†. Cell Reports (2013). PDF
Mouse p202 containing two hemopoietic expression, interferon inducibility, nuclear localization (HIN) domains antagonizes AIM2 inflammasome signaling and potentially modifies lupus susceptibility. We found that only HIN1 of p202 binds double-stranded DNA (dsDNA), while HIN2 forms a homotetramer. Read More
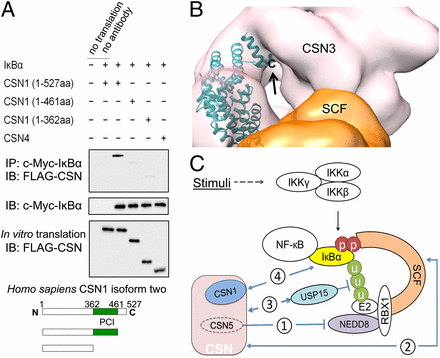
Crystal structure and versatile functional roles of the COP9 signalosome subunit 1
Lee JH*, Yi L*, Li J, Schweitzer K, Borgmann M, Naumann M, Wu H†. Proc Natl Acad Sci U S A (2013). PDF
The constitutive photomorphogenesis 9 (COP9) signalosome (CSN) plays key roles in many biological processes, such as repression of photomorphogenesis in plants and protein subcellular localization, DNA-damage response, and NF-κB activation in mammals. It is an evolutionarily conserved eight-protein complex with subunits CSN1 to CSN8 named following the descending order of molecular weights. Here, we report the crystal structure of the largest CSN subunit, CSN1 from Arabidopsis thaliana (atCSN1), which belongs... Read More

Higher-Order Assemblies in a New Paradigm of Signal Transduction
Recent studies have revealed that multiple intracellular signaling proteins may assemble into structured, yet sometimes infinite, higher-order signaling machines for transmission of receptor activation information to cellular responses. These studies advance our understanding of cell signaling and implicate new molecular mechanisms in proximity-driven enzyme activation, threshold behavior, signal amplification, reduction of biological noise, and temporal and spatial control of signal transduction... Read More

MALT1 Small Molecule Inhibitors Specifically Suppress ABC-DLBCL In Vitro and In Vivo.
Fontan L*, Yang C*, Kabaleeswaran V, Volpon L, Osborne MJ, Beltran E, Garcia M, Cerchietti L, Shaknovich R, Yang SN, Fang F, Gascoyne RD, Martinez-Climent JA, Glickman JF, Borden K, Wu H†, Melnick A†. Cancer Cell (2012). PDF
MALT1 cleavage activity is linked to the pathogenesis of activated B cell-like diffuse large B cell lymphoma
(ABC-DLBCL), a chemoresistant form of DLBCL. We developed a MALT1 activity assay and identified chemically diverse MALT1 inhibitors. A selected lead compound, MI-2, featured direct binding to MALT1... Read More

Cyclic di-GMP Sensing via the Innate Immune Signaling Protein STING
Detection of foreign materials is the first step of successful immune responses. Stimulator of interferon genes (STING) was shown to directly bind cyclic diguanylate monophosphate (c-di-GMP), a bacterial second messenger, and to elicit strong interferon responses. Here we elucidate the structural features of the cytosolic c-di-GMP binding domain (CBD) of STING and its complex with c-diGMP. The CBD exhibits an a + b fold and is a dimer in the crystal and in solution. Read More
.jpg)
The RIP1/RIP3 Necrosome Forms a Functional Amyloid Signaling Complex
Required for Programmed Necrosis
Li J, McQuade T, Siemer AB, Napetschnig J, Moriwaki K, Hsiao YS, Damko E, Moquin D, Walz T, McDermott A, Chan FK, Wu H†. Cell (2012). PDF
RIP1 and RIP3 kinases are central players in TNF-induced programmed necrosis. Here, we report that the RIP homotypic interaction motifs (RHIMs) of RIP1 and RIP3 mediate the assembly of heterodimeric filamentous structures. The fibrils exhibit classical characteristics of b-amyloids, as shown by... Read More
Other Publications (2012-2016)
* = co-first author, † = corresponding author
Janssen E*,Tohme M*, Hedayat M*, Leick M*,Kumari S, Ramesh N, Massaad M J, Ullas S, Azcutia V, Goodnow C C, Randall K L, Qiao Q, Wu H, Al-Herz W, Cox D, Hartwig J, Irvine D J, Luscinskas F W, and Geha RS† (2016). A DOCK8-WIP-WASp complex links T cell receptors to the actin cytoskeleton. The Journal of Clinical Investigation. 126(10)3837-3851
Zanoni I, Tan Y, Di Gioia M, Broggi A, Ruan J, Shi J, Donado CA, Shao F, Wu H, Springstead JR, Kagan JC† (2016). An endogenous caspase-11 ligand elicits interleukin-1release from living dendritic cells. Science. 352:1232-6
Myer PA, Socias S, Key J, Ransey E, Tjon EC, Buschiazzo A, Lei M, Botka C, Withrow J, Neau D, Rajashankar K, Anderson KS, Baxter RH, Blacklow SC, Boggon TJ, Bonvin A, Borek D, Brett TJ, Caflisch A, Chung-I C, Chazin WJ, Corbett KD, Cosgrove MS, Crosson S, Dhe-Paganon S, Cera ED, Drennan CL, Eck MJ, Eichman BF, Fan QR, Ferré-D’Amaré AR, Fromme JC, Garcia KC, Gaudet R, Gong P, Harrison SC, Heldwein EE, Jia Z, Keenan RJ, Kruse AC, Kvansakul M, McLellan JS, Modis Y, Nam Y, Otwinowski Z, Pai EF, Pereira PJB, Petosa C, Raman CS, Rapoport TA, Roll-Mecak A, Rosen MK, Rudenko G, Schlessinger J, Schwartz TU, Shamoo Y, Sondermann H, Tao YJ, Tolia NH, Tsodikov OV, Westover KD, Wu H, Foster I, Fraser JS, Maia FR, Gonen T, Kirchhausen T, Diederichs K, Crosas M, Sliz P†(2016). Data publication with the structural biology data grid supports live analysis. Nature Communications. 7;7:10882
Qiao Q, Wu H† (2015). Supramolecular organizing centers (SMOCs) as signaling machines in innate immune activation. Sci China Life Sci. 58:1067-72
Vajjhala P, Lu A, Brown D, Pang S, Sagulenko V, Sester D, Cridland S, Hill J, Schroder K, Stow J, Wu H†, Stacey K†. (2015). The inflammasome adaptor ASC induces procaspase-8 death effector domain filaments. J Biol Chem. 290:29217-30
Lu A, Wu H† (2015). Structural Mechanism of Inflammasome Assembly. FEBS J. 282:435-444
Wu H*, Yokoyama S*, Lugar K*, Doudna J*, Wilson I*, Xu RM*. (2015). What’s your favorite crystal structure? Cell. 159:967-968
Lu A*, Li Y*, Qian Y, Ruan J, Yu X, Egelman E, Wu H†. (2015). Plasticity in PYD assembly revealed by cryo-EM structure of the PYD filament of AIM2. Cell Discovery. 1, 15013
Onizawa M*, Oshima S*, Schulze-Topphoff U*, Oses-Prieto JA, Lu T, Tavares R, Prodhomme T, Duong B, Whang MI, Advincula R, Agelidis A, Barrera J, Wu H, Burlingame A, Malynn BA, Zamvil SS, Ma A†. (2015). The ubiquitin-modifying enzyme A20 restricts ubiquitination of the kinase RIP3 and protects cells from necroptosis. Nature Immunology. 16:618-627
Wei S, Kozono S, Kats L, Nechama M, Li W, Guarnerio J, Luo M, You MH, Yao Y, Kondo A, Hu H, Bozkurt G, Moerke NJ, Cao S, Reschke M, Chen CH, Rego EM, Lo-Coco F, Cantley LC, Lee TH, Wu H, Zhang Y, Pandolfi PP, Zhou XZ†, Lu KP† (2015). Active Pin1 is a key target of all-trans retinoic acid in acute promyelocytic leukemia and breast cancer. Nat Med. 21:457-466
Hauenstein A*, Zhang L*, Wu H†. (2015). The hierarchical structural architecture of inflammasomes, supramolecular inflammatory machines. Curr Opin Struct Biol. 31:75-83
Li L*†, Yin Q, Kuss P, Maliga Z, Millán JL, Wu H, Mitchison T*. (2014). Hydrolysis of 2'3'-cGAMP by ENPP1 and design of nonhydrolyzable analogs. Nat Chem Biol. 10:1043-8
Jang TH, Zheng C, Li J, Richards C, Hsiao YS, Walz T, Wu H, Park HH†. (2014). Structural Study of the RIPosptosome core reveals a helical assembly for kinase recruitment. Biochemistry. 53:5424-31
Ru H*, Zhang P*, Wu H†. (2014). Promiscuity is not Always Bad. Mol Cell. 2:208-9
Furman RR*, Cheng S*, Lu P*, Setty M, Perez AR, Guo A, Racchumi J, Xu G, Wu H, Ma J, Steggerda SM, Coleman M, Leslie C, Wang YL†. (2014). Ibrutinib Resistance in Chronic Lymphocytic Leukemia. N Engl J Med. 24:2352-4
Lu A, Kavaleeswaran V, Fu T, Magupalli VG, Wu H†. (2014). Crystal Structure of the F27G AIM2 PYD Mutant and Similarities of Its Self-Association to DED/DED Interactions. J Mol Biol. 7:1420-7
Yang C†, David L, Qiao Q, Damko E, Wu H. (2013). The CBM Signalosome: Potential Therapeutic Target for Aggressive Lymphoma? Cytokine Growth Factor Rev. 2:175-183
Kim S*, Li L*, Maliga Z, Yin Q, Wu H, Mitchison T†. (2013). Anticancer Flavonoids Are Mouse-Selective STING Agonists. ACS Chemical Biology. 8:1396-1401
Li J*, Yin Q*, Wu H†. (2013). Structural Basis of Signal Transduction in the TNF Receptor Superfamily. Advanced Immunology. 119:135-53
Napteschnig J, Wu H†. (2013). Molecular Basis of NF-kB signaling. Annual Reviews in Biophysics. 42:443-68
Lu TT, Onizawa M, Hammer GE, Turer EE, Yin Q, Damko E, Agelidis A, Shifrin N, Advincula R, Barrera J, Malynn BA, Wu H, Ma A†. (2013). Dimerization and Ubiquitin Mediated Recruitment of A20, a Complex Deubiquitinating Enzyme. Immunity. 38:1-10
Ferrao R, Li J, Bergamin E, Wu H†. (2012). Structural insights into the assembly of large oligomeric signalosomes in the Toll-like receptor-interleukin-1 receptor superfamily. Sci Signal. 5:re3
Ferrao R, Wu H†. (2012). Helical assembly in the death domain (DD) superfamily. Curr Opin Struct Biol. 22:241-7
Feldman T, Kabaleeswaran V, Jang SB, Antczak C, Djaballah H, Wu H, Jiang X†. (2012). A class of allosteric caspase inhibitors identified by high-throughput screening. Mol Cell. 47:585-95

